13 releases
| 0.3.9 | May 10, 2023 |
|---|---|
| 0.3.8 | Mar 23, 2023 |
| 0.3.2 | Jan 3, 2023 |
| 0.3.1 | Nov 29, 2022 |
| 0.1.1 | Nov 22, 2022 |
#58 in Biology
164 downloads per month
52KB
1K
SLoC
rnapkin: drawing RNA secondary structure with style
Usage
rnapkin accepts a file containing secondary structure and optionally sequence and a name. For example you could have this marvelous RNA molecule sitting peacefully in a file called "guaniners"
>fantastic guanine riboswitch
AAUAUAAUAGGAACACUCAUAUAAUCGCGUGGAUAUGGCACGCAAGUUUCUACCGGGCAC
..........(..(.((((.((((..(((((.......)))))..........((((((.
CGUAAAUGUCCGACUAUGGGUGAGCAAUGGAACCGCACGUGUACGGUUUUUUGUGAUAUC
......)))))).....((((((((((((((((((........))))))...........
AGCAUUGCUUGCUCUUUAUUUGAGCGGGCAAUGCUUUUUUUA
..)))))))))))).)))).)))).)..).............
Then, if you wish to visualize it, you could invoke rnapkin thus:
rnapkin guaniners
Surely rnapkin would respond with the name of a file it has just drawn to:
fantastic_guanine_riboswitch.svg
And this scalable vector graphic would be produced:
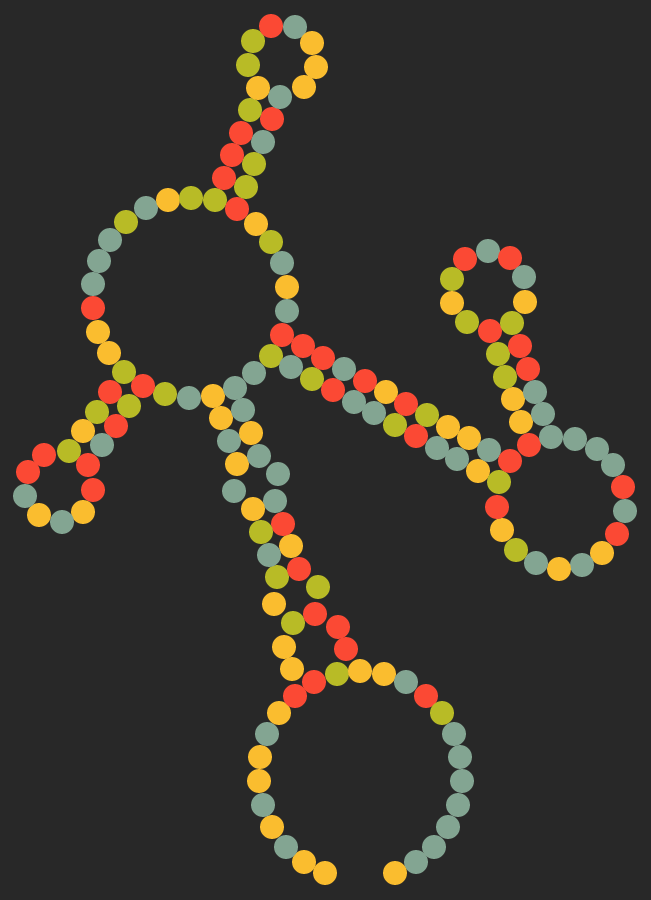
I happen to quite enjoy the outcome, so I would say:
that's pretty neat
Your mileage may vary though.
Rotating and flipping
If you'd like to see this or any other RNA molecule upside-down, tilted or what have you, there are some options listed below that you can use and combine:
-a / --angle <DEGREES> | starting Angle / boils down to clockwise rotation
--mx | Mirror along X axis / aka vertical flip
--my | Mirror along Y axis / aka horizontal flip
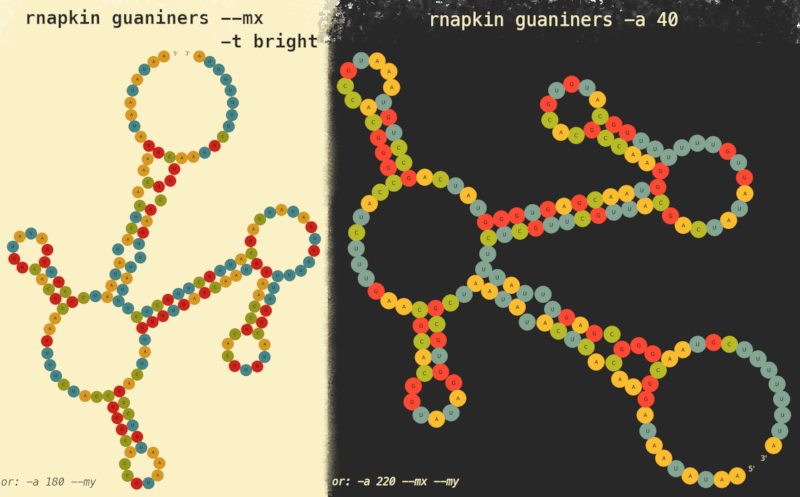

color themes can be changed by -t option as demonstrated; a config file allowing to define custom color themes is planned though unimplemented!()
Installing
I plan to offer precompiled binaries but for now you'll need rust. Easiest way to acquire rust is via rustup 🦀
Anywhere
cargo install rnapkin
WSL
Fontconfig is the default Fontmanagement utility on Linux and Unix but WSL may not have them installed;
sudo apt-get install libfontconfig libfontconfig1-dev
cargo install rnapkin
Input
input can be served to rnapkin as a file or be piped in:
rnapkin cmolecule.fa -a 20 -o crab
echo ".......(((((......))))).....(((((......)))))......." | rnapkin -a 20 -o crab
input is quite flexible; it should contain secondary_structure and optionally name and sequence. Name has to start with ">" and can be overwritten with -o flag which has priority. Here are some variations of valid input files:
simple one
# you can add .png to the name to request png instead of svg
@ the same of course can be achieved with -o flag.
* this is a comment btw: any symbol other than ">.()" works but prefer "#"
>simple molecule.png
((((((((((..((((((.........))))))......).((((((.......))))))..)))))))))
CGCUUCAUAUAAUCCUAAUGAUAUGGUUUGGGAGUUUCUACCAAGAGCCUUAAACUCUUGAUUAUGAAGUG
Highlighting!
There are 9 available colors denoted by 1-9, while 0 means None. For example consider input below representing SAM riboswitch in the OFF conformation according to smFRET study by Manz et al. 2017. By using numbers in the input we can mark aptamer forming helices P1, P2, P3, P4 #2-#5 and the TERMINATOR #1.
> offsam
0000022222222223333333333333333333333333333333333444444444444444444444444444444
AUAUCCGUUCUUAUCAAGAGAAGCAGAGGGACUGGCCCGACGAUGCUUCAGCAACCAGUGUAAUGGCGAUCAGCCAUGA
.......((((((((....(((((...(((.....)))......)))))(((..(((((...(((((.....))))).)
4444444444555555555555555555555555555522222222222211111111111111111111111111111
CUAAGGUGCUAAAUCCAGCAAGCUCGAACAGCUUGGAAGAUAAGAAGAGACAAAAUCACUGACAAAGUCUUCUUCUUAA
))..)).)))........((((((.....))))))...)))))))).................((((((.((((...))
111111111111
GAGGACUUUUUU
)).))))))...
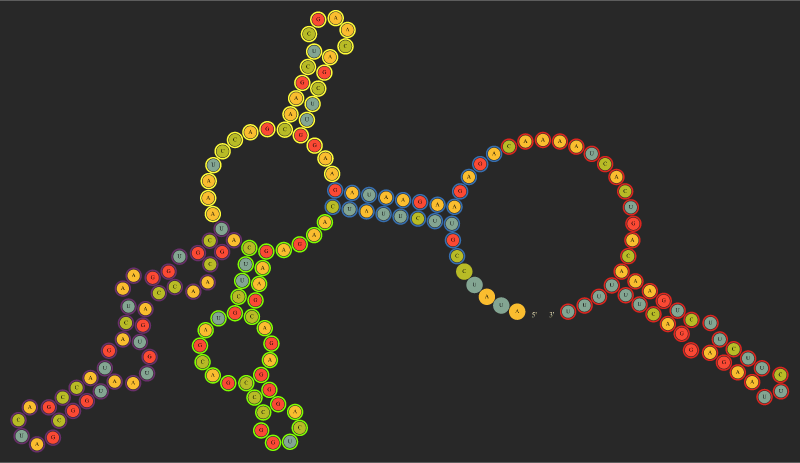
only secondary structure
.........(((..((((((...((((((((.....((((((((((...)))))).....
(((((((...))))))).))))(((.....)))...)))).)))).))))))..)))..(
(((.(((((..(((......))).)))))..))))(((((((((((((....))))))))
))))).....
multiline
sequence and secondary structure can be separate, mixed and aligned, everything should work.
DIY
using -p / --points flag you can make rnapkin print calculated coordinates of nucleotide bubbles (with 0.5 unit radius). You can then plot it yourself if you need to do something specific;
If you happen to clone the repository, there is an example python script using matplotlib that you can pipe the input to.
cargo run -- atelier/example_inputs/guaniners -p | atelier/plot.py
You can also combine -p flag with --mx --my and -a
rnapkin name
The wordsmithing proccess was arduous. It involved googling "words starting with na" and looking for anything drawing related. Once the word was found, unparalled strength was employed to slap it on top of "rna" ultimately creating this glorious amalgamation.
why it kinda makes sense:
You ever heard of all those physicists, mathematicians and the like, scribbling formulas on the
back of a napkin or a book margin? There is even a wikipedia page about it.
It doesn't take much mental gymnastic to imagine a biologist frantically scrambling together rna structure on a napkin. I am currently working on baiting my biologist friend into heated rna debate while in close proximity to abundant napkin source in order to produce a proof of concept.
Dependencies
~5.5–8.5MB
~141K SLoC