2 unstable releases
| 0.2.0 | Feb 17, 2022 |
|---|---|
| 0.1.0 | Feb 16, 2022 |
#356 in Science
195KB
580 lines
proteinogenic 
Chemical structure generation for protein sequences as SMILES string.
🔌 Usage
This crate builds on top of purr, a crate providing
primitives for reading and writing SMILES.
Use the AminoAcid enum to encode the sequence residues, and build a SMILES
string with proteinogenic::smiles. For example with divergicin 750:
extern crate proteinogenic;
let residues = "KGILGKLGVVQAGVDFVSGVWAGIKQSAKDHPNA"
.chars()
.map(proteinogenic::AminoAcid::from_char)
.map(Result::unwrap);
let s = proteinogenic::smiles(residues)
.expect("failed to generate SMILES string");
Additional modifications can be carried out by using a Peptide struct to
configure the rendering of the peptide. So far, disulfide bonds as well as
lanthionine bridges are supported, as well as head-to-tail cyclization.
For instance. we can generate the SMILES string of a
cyclotide such as
kalata B1:
extern crate proteinogenic;
let residues = "GLPVCGETCVGGTCNTPGCTCSWPVCTRN"
.chars()
.map(proteinogenic::AminoAcid::from_char)
.map(Result::unwrap);
let mut p = proteinogenic::Protein::new(residues);
p.cyclization(proteinogenic::Cyclization::HeadToTail);
p.cross_link(proteinogenic::CrossLink::Cystine(5, 19)).unwrap();
p.cross_link(proteinogenic::CrossLink::Cystine(9, 21)).unwrap();
p.cross_link(proteinogenic::CrossLink::Cystine(14, 26)).unwrap();
let s = p.smiles()
.expect("failed to generate SMILES string");
This SMILES string can be used in conjunction with other cheminformatics toolkits, for instance OpenBabel which can generate a PNG figure:
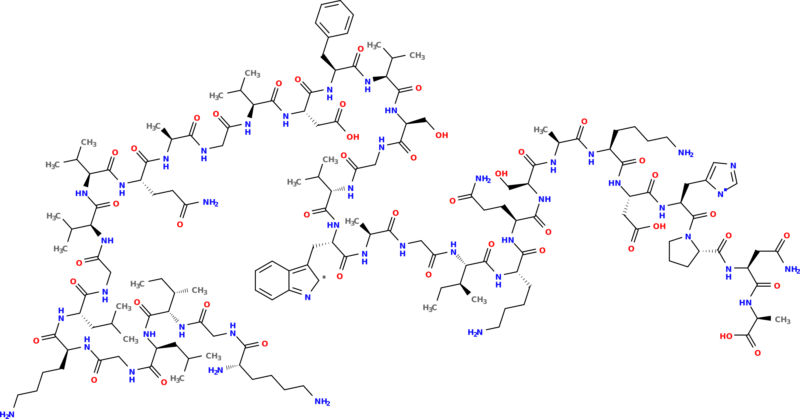
Note that proteinogenic is not limited to building a SMILES string; it can
actually use any purr::walk::Follower
implementor to generate an in-memory representation of a protein formula. If
your code is already compatible with purr, then you'll be able to use
protein sequences quite easily.
extern crate proteinogenic;
extern crate purr;
let sequence = "KGILGKLGVVQAGVDFVSGVWAGIKQSAKDHPNA";
let residues = sequence.chars()
.map(proteinogenic::AminoAcid::from_char)
.map(Result::unwrap);
let mut builder = purr::graph::Builder::new();
proteinogenic::visit(residues, &mut builder);
builder.build()
.expect("failed to create a graph representation");
The API is not yet stable, and may change to follow changes introduced by
purr or to improve the interface ergonomics.
💭 Feedback
⚠️ Issue Tracker
Found a bug ? Have an enhancement request ? Head over to the GitHub issue tracker if you need to report or ask something. If you are filing in on a bug, please include as much information as you can about the issue, and try to recreate the same bug in a simple, easily reproducible situation.
📋 Changelog
This project adheres to Semantic Versioning and provides a changelog in the Keep a Changelog format.
🔍 See Also
If you're a bioinformatician and a Rustacean, you may be interested in these other libraries:
uniprot.rs: Rust data structures for the UniProtKB databases.obofoundry.rs: Rust data structures for the OBO Foundry.fastobo: Rust parser and abstract syntax tree for Open Biomedical Ontologies.pubchem.rs: Rust data structures and API client for the PubChem API.
📜 License
This library is provided under the open-source MIT license.
This project was developed by Martin Larralde during his PhD project at the European Molecular Biology Laboratory in the Zeller team.